 
Consider lowering the sample protein concentration.Samples that are too concentrated or are poorly denatured will not separate cleanly-this is important for the appearance of crisp bands. The primary antibody may just be of lower quality for your purposes, and another company’s (using a different epitope or purification method) may work better.Find the optimal concentration with a titration blot-using too much primary can lead to extraneous binding, and too little can lead to problems with no signal at all.If you are using film, you may consider switching to an imager. Experiment with different imaging protocols and contrast settings to find which can produce a clean signal with minimal exposure time.Test the membrane (and substrate)-add your imaging substrate to an empty, non-treated membrane to ensure you get no signal when there is no secondary bound.1:1000 dilution is pretty standard, but 1:5000 or even 1:10000 may be indicated. Lower the concentration of your secondary antibody.Don’t exceed the recommended incubation times, both for secondary antibody and your imaging agent!.Here’s where the signal is made-literally. Increasing the speed/vigor of the shaker, or washing for a greater amount of time.5 rounds of 6 minutes instead of 3 rounds of 10) Over-washing can diminish the signal of interest, but this isn’t your problem if you have high background noise. The whole purpose of washing is to clear the membrane of non-specific, weak interactions that eventually result in background noise. Comparing with a different blocking buffer entirely.Using a higher the protein concentration in your buffer.Increasing the blocking exposure time and/or temperature at which you block.If you’re having trouble with non-specific binding, consider: 
This is the most important step of the blot-if you don’t block the unoccupied sites on the membrane, the antibodies will bind directly to the membrane. Let’s go through some ways to sharpen up your blot, in order of relative importance. But alternatively, what do you do when too much background is the problem? You may have beautiful bands of interest-but if there is a bunch of non-specific binding, your quantification and data reliability will suffer. After discarding the last PBST wash, add approximately 1 milliliter of horseradish peroxide substrate to the membrane, and detect the chemoluminescent signal.In the previous installment of this series on western blotting, we addressed potential sources of error when your final product is completely bare. Wash the membrane three times with PBST again.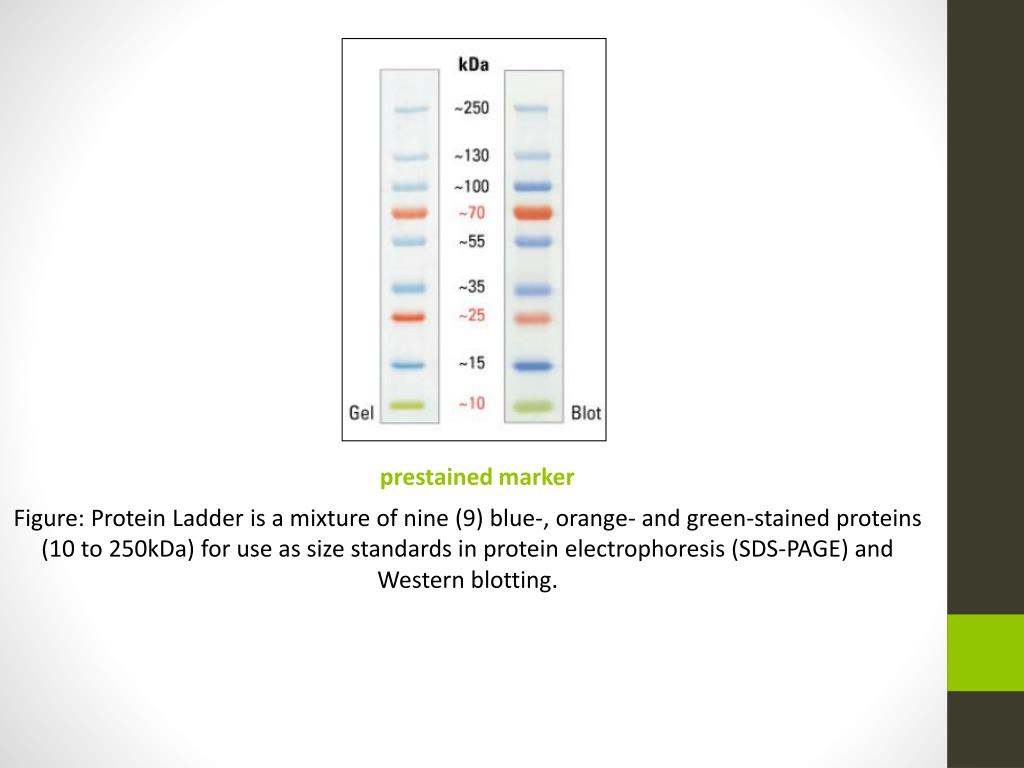  
The next day, wash the membrane with three 5-minute PBST washes, then, incubate the membrane with 20 milliliters of secondary antibody with gentle shaking for 1 hour at room temperature. To detect the proteins, add the first primary antibody at the indicated dilution to the membrane, and incubate it overnight at 4 degrees Celsius with gentle agitation. Then, wash the membrane three times for 5 minutes with PBST. After the transfer is complete, place the blotted membrane in a box, and incubate it for an hour in blocking solution, under gentle agitation. Load the lysates, immunoprecipitants, and a protein standard onto a 12.5% SDS gel, and run with a constant voltage of 80 volts.Īfter the gel run is complete, transfer the proteins from the SDS gel to a nitrocellulose membrane. Simultaneously, heat the lysate controls at 95 degrees Celsius for 5 minutes. To perform the western blot, add 20 microliters of 4X loading buffer to the beads, and heat at 95 degrees Celsius for 10 minutes. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
